|
Cholesterol study suggests new diagnostic, treatment approach for prostate cancer
Date:March 4, 2014 Source: Purdue University Summary: A link between prostate cancer aggressiveness and the accumulation of a compound produced when cholesterol is metabolized in cells has been discovered, findings that could bring new diagnostic and treatment methods. Findings also suggest that a class of drugs previously developed to treat atherosclerosis might be repurposed for treatment of advanced prostate cancer. The research involved analysis of clinical samples harvested from prostate cancer patients, specialized cell lines and mice. Full Story: Researchers have discovered a link between prostate cancer aggressiveness and the accumulation of a compound produced when cholesterol is metabolized in cells, findings that could bring new diagnostic and treatment methods. Findings also suggest that a class of drugs previously developed to treat atherosclerosis might be repurposed for treatment of advanced prostate cancer. The research showed depletion of the compound cholesteryl ester significantly reduced prostate cancer cell proliferation, impaired its ability to invade a laboratory tissue culture and suppressed tumor growth in mice. "Our study provides an avenue towards diagnosis of aggressive prostate cancer. Moreover, we showed that depleting cholesteryl ester significantly impairs prostate cancer aggressiveness," said Ji-Xin Cheng, a professor in Purdue University's Weldon School of Biomedical Engineering and Department of Chemistry. The research involved analysis of clinical samples harvested from prostate cancer patients, specialized cell lines and mice. Findings are detailed in a research paper appearing in the journal Cell Metabolism. The paper was authored by researchers associated with Purdue's Center for Cancer Research and the Indiana University Melvin and Bren Simon Cancer Center at Indiana University School of Medicine. "Prostate cancer is the second-leading cause of cancer-related mortality in American men. Our finding offers a biological foundation that supports the beneficial effect of cholesterol-lowering drugs. Second, our study heralds the potential of using cholesteryl ester as a therapeutic target for advanced prostate cancer," said study co-author Timothy Ratliff, the Robert Wallace Miller Director of Purdue's Center for Cancer Research. "These results together suggest that cholesteryl ester accumulation might be used for more accurate prediction of prostate cancer aggressiveness, if validated through further examination of a large number of tissue biopsies and correlation assessment of cholesteryl ester levels and clinical outcomes of patients." A critical focus of the research is the analysis of individual lipid droplets inside single cells. Purdue researchers have developed an analytical tool called Raman spectromicroscopy that allows compositional analysis of single lipid droplets in living cells and mice. "It is conceivable that cancer cells require reservoirs for lipids, namely lipid droplets. However, our imaging data revealed an unexpected, aberrant accumulation of esterified cholesterol in lipid droplets of high-grade prostate cancer and metastases," Cheng said. The researchers learned that cholesteryl ester accumulation, which occurs only in advanced prostate cancer and its metastasis, results from the loss of a tumor-suppressing gene called PTEN and the activation of an intracellular metabolic pathway promoting tumor growth. "These findings improve current understanding of the role of cholesterol in cancer and also suggest new opportunities for the diagnosis and treatment of aggressive prostate cancer. We have been pleased to be able to collaborate with Dr. Cheng on his important research" said Michael Koch, John P. Donohue Professor of Urology and chair of the Department of Urology at IU School of Medicine. Findings show the drugs avasimibe and Sandoz 58-035 reduced the accumulation of cholesteryl ester and significantly hindered advanced prostate cancer growth in laboratory cell cultures and xenograft mouse models. These drugs did not show toxicity to animals. "We note that avasimibe, Sandoz 58-035 and a class of similar drugs were developed to treat atherosclerosis, but the clinical trials were halted due to the lack of effectiveness in reducing plaque size," Cheng said. "The present study highlights a novel use of these drugs to treat advanced prostate cancer." Story Source: The above story is based on materials provided by Purdue University. The original article was written by Emil Venere. Note: Materials may be edited for content and length. 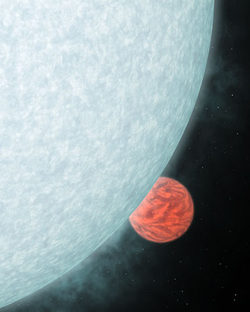
'Dimer Molecules' Aid Study of Exoplanet Pressure, Hunt for Life
Date:March 4, 2014 Source: University of Washington Summary: Astronomers have developed a new method of gauging the atmospheric pressure of exoplanets, or worlds beyond the solar system, by looking for a certain type of molecule. And if there is life out in space, scientists may one day use this same technique to detect its biosignature — the telltale chemical signs of its presence — in the atmosphere of an alien world. Astronomers at the University of Washington have developed a new method of gauging the atmospheric pressure of exoplanets, or worlds beyond the solar system, by looking for a certain type of molecule. And if there is life out in space, scientists may one day use this same technique to detect its biosignature -- the telltale chemical signs of its presence -- in the atmosphere of an alien world. Understanding atmospheric pressure is key to knowing if conditions at the surface of a terrestrial, or rocky, exoplanet might allow liquid water, thus giving life a chance. The method, devised by Amit Misra, a UW astronomy doctoral student, and co-authors, involves computer simulations of the chemistry of Earth's own atmosphere that isolate what are called "dimer molecules" -- pairs of molecules that tend to form at high pressures and densities in a planet's atmosphere. There are many types of dimer molecules but this research focused only on those of oxygen. Misra and team ran simulations testing the spectrum of light in various wavelengths. Dimer molecules absorb light in a distinctive pattern, and the rate at which they form is sensitive to the pressure, or density, in the planet's atmosphere. "So the idea is that if we were able to do this for another planet, we could look for this characteristic pattern of absorption from dimer molecules to identify them," Misra said. The presence of such molecules, he said, likely means the planet has at least one-quarter to one-third the pressure of Earth's atmosphere. Powerful telescopes soon to come online, such as the James Webb Space Telescope, scheduled for launch in 2018, may enable astronomers to use this method on distant exoplanets. With such enhanced tools, Misra said, astronomers might detect dimer molecules in actual exoplanet atmospheres, leading to a clear understanding of the planet's atmosphere. This research may also play a part in the greatest astronomical quest of all -- the ongoing search for life in the cosmos. That's because the team realized along the way that oxygen dimer molecules are often more detectable in an atmosphere than other markers of oxygen. That's important from a biological standpoint, Misra said. "It's tied to photosynthesis, and we have pretty good evidence that it's hard to get a lot of oxygen in an atmosphere unless you have algae or plants that are producing it at a regular rate. "So if we find a good target planet, and you could detect these dimer molecules -- which might be possible within the next 10 to 15 years -- that would not only tell you something about pressure, but actually tell you that there's life on that planet." Misra's UW co-author is Victoria Meadows, professor of astronomy; other co-authors are Mark Claire of Scotland's University of St. Andrews and Dave Crisp of NASA's Jet Propulsion Laboratory in Pasadena, Calif. The team's paper was published in the February issue of the journal Astrobiology. The research was performed through the UW-based Virtual Planetary Laboratory and funded by NASA (Grant NNH05ZDA001C), as well as a grant from Advancing Science in America, Seattle chapter. Story Source: The above story is based on materials provided by University of Washington. The original article was written by Peter Kelley. Note: Materials may be edited for content and length. |
Imprint of chemotherapy linked to inflammation in breast cancer survivors
Date: March 4, 2014 Source: Emory Health Sciences Summary: Chemotherapy can leave a long-lasting epigenetic imprint in the DNA of breast cancer patients' blood cells. That imprint is associated with biological signs of inflammation up to six months after the completion of treatment, and many breast cancer survivors experience fatigue and other debilitating symptoms that persist months to years after their course of treatment has ended. Now researchers have found clues that may explain how these symptoms can linger. Full Story: ny breast cancer survivors experience fatigue and other debilitating symptoms that persist months to years after their course of treatment has ended. Now researchers at the Winship Cancer Institute of Emory University have found clues that may explain how these symptoms can linger. Chemotherapy, one of the major treatments for breast cancer, can leave a long-lasting epigenetic imprint in the DNA of breast cancer patients' blood cells. That imprint is associated with biological signs of inflammation up to six months after the completion of treatment. Inflammation in turn is believed to cause symptoms like fatigue. The findings are published online in the journal Brain, Behavior and Immunity. "Chemotherapy is a life-saving intervention, but for some women it comes at a cost," says Mylin Torres, MD, assistant professor of radiation oncology at Emory University School of Medicine and Winship Cancer Institute. "These results are the first to suggest a biological mechanism by which treatment-related side effects can persist long after treatment completion in women with breast cancer." That achy, tired feeling that comes from the flu is caused by inflammation, the body's natural response to infection or a wound -- but it usually disappears once an illness is over. In patients being treated for cancer, fatigue has also been linked to inflammation and up to 30 percent of breast cancer survivors experience persistent fatigue long after treatment has ended. To investigate the biology behind persistent cancer-related inflammation, Torres teamed up with Andrew Miller, MD, professor of psychiatry and behavioral sciences and director of psychiatric oncology at Winship Cancer Institute. First co-authors of the paper are Alicia Smith, PhD, assistant professor of psychiatry and behavioral sciences, and Karen Conneely, PhD, assistant professor of human genetics. In a previously published study led by Torres and Miller, chemotherapy was associated with increased markers of inflammation in the blood which correlated with fatigue. "We had found that increased fatigue was similar no matter whether patients received chemotherapy before or after surgery," Torres says, "indicating that the timing of chemotherapy was less important than whether you got it in the first place." In the current study, all the women went through partial mastectomy surgery. Some received varying forms of chemotherapy, and all received radiation at the end. They found that women who had been treated with chemotherapy exhibited changes in the methylation of the DNA in their white blood cells. Some of these changes were still present six months after radiation. Methylation is an epigenetic alteration in DNA, which does not change the A/C/G/T "letter" information in the DNA but does change how that information is read by the cell, influencing whether a gene is turned on or off. The researchers scanned hundreds of thousands of potential sites of methylation; only eight sites were reliably altered in women who received chemotherapy, and changes at half of those sites were visible six months later. The biology connecting this handful of sites to inflammation remains unclear; the eight sites were not in genes that encode inflammatory signaling proteins secreted into the blood, for example. However, the methylation changes did correlate with increased levels of two inflammatory signaling proteins, IL-6 and sTNFR2, that have been associated with fatigue in breast cancer survivors. Many chemotherapeutic agents are effective precisely because they damage DNA in cancer cells. While it was well known that chemotherapy induces epigenetic changes in cancer cells, this is the first study to identify epigenetic changes induced by chemotherapy for breast cancer in non-cancerous cells of the blood. The authors hypothesize that chemotherapy may directly alter methylation status in the blood cells, or it may be a result of the inflammatory response to chemotherapy-related tissue injury. "It may be something about the intensity or the repetitive nature of chemotherapy that makes it qualitatively different from acute inflammation," Miller says. "The more we know about this imprinting process, the better chance we have of getting to new therapies for chronic treatment-related problems, such as fatigue, in breast cancer survivors." Story Source: The above story is based on materials provided by Emory Health Sciences. Note: Materials may be edited for content and length. 
First look at how Staphylococcus cells adhere to nanostructures could help fight infections
Date:March 4, 2014 Source: DOE/Lawrence Berkeley National Laboratory Summary: A team of researchers has explored, for the first time, how individual Staphylococcus cells glom onto metallic nanostructures of various shapes and sizes that are not much bigger than the cells themselves. Their work could lead to a more nuanced understanding of what makes a surface less inviting to bacteria. A Staph infection can't start unless Staphylococcus cells first cling to a surface, which is why scientists are hard at work exploring bacteria-resistant materials as a line of defense. This scanning electron microscopy image reveals how Staphylococcus Aureus cells physically interact with a nanostructure. A bacterial cell (blue) is embedded inside the hollow nanopillar's hole and several cells cling to the nanopillar's curved walls. Credit: Mofrad lab and the Nanomechanics Research Institute [Click to enlarge image] The bacterium Staphylococcus Aureus (S. aureus) is a common source of infections that occur after surgeries involving prosthetic joints and artificial heart valves. The grape-shaped microorganism adheres to medical equipment, and if it gets inside the body, it can cause a serious and even life-threatening illness called a Staph infection. The recent discovery of drug-resistant strains of S. aureus makes matters even worse. A Staph infection can't start unless Staphylococcus cells first cling to a surface, however, which is why scientists are hard at work exploring bacteria-resistant materials as a line of defense. This research has now gone nanoscale, thanks to a team of researchers led by Berkeley Lab scientists. They investigated, for the first time, how individual S. aureuscells glom onto metallic nanostructures of various shapes and sizes that are not much bigger than the cells themselves. They found that bacterial adhesion and survival rates vary depending on the nanostructure's shape. Their work could lead to a more nuanced understanding of what makes a surface less inviting to bacteria. "By understanding the preferences of bacteria during adhesion, medical implant devices can be fabricated to contain surface features immune to bacteria adhesion, without the requirement of any chemical modifications," says Mohammad Mofrad, a faculty scientist in Berkeley Lab's Physical Biosciences Division and a professor of Bioengineering and Mechanical Engineering at UC Berkeley. Mofrad conducted the research with the Physical Biosciences Division's Zeinab Jahed, the lead author of the study and a graduate student in Mofrad's UC Berkeley Molecular Cell Biomechanics Laboratory, in collaboration with scientists from Canada's University of Waterloo. Their research was recently published online in the journal Biomaterials. The scientists first used electron beam lithographic and electroplating techniques to fabricate nickel nanostructures of various shapes, including solid pillars, hollowed-out pillars, c-shaped pillars, and x-shaped columns. These features have outer diameters as small as 220 nanometers. They also created mushroom-shaped nanostructures with tiny stems and large overhangs. They introduced S. aureus cells to these structures, gave the cells time to stick, and then rinsed the structures with deionized water to remove all but the most solidly bound bacteria. Scanning electron microscopy revealed which shapes are the most effective at inhibiting bacterial adhesion. The scientists observed higher bacteria survival rates on the tubular-shaped pillars, where individual cells were partially embedded into the holes. In contrast, pillars with no holes had the lowest survival rates. The scientists also found that S. aureus cells can adhere to a wide range of surfaces. The cells not only adhere to horizontal surfaces, as expected, but to highly curved features, such as the sidewalls of pillars. The cells can also suspend from the overhangs of mushroom-shaped nanostructures. "The bacteria seem to sense the nanotopography of the surface and form stronger adhesions on specific nanostructures," says Jahed. Story Source: The above story is based on materials provided by DOE/Lawrence Berkeley National Laboratory. The original article was written by Dan Krotz. Note: Materials may be edited for content and length. |

